Cloudknot:
Scaling your existing (Python) code in the (AWS) cloud
The University of Washington
Follow along at http://arokem.github.io/2020-OHBM-cloudknot/

We love working in our Python environment
We also like to use cloud computing
Pros:
- Linear scalability
- Elasticity
- Ability to handle large datasets
Cons:
- Learning curve
- Complexity
- Cost
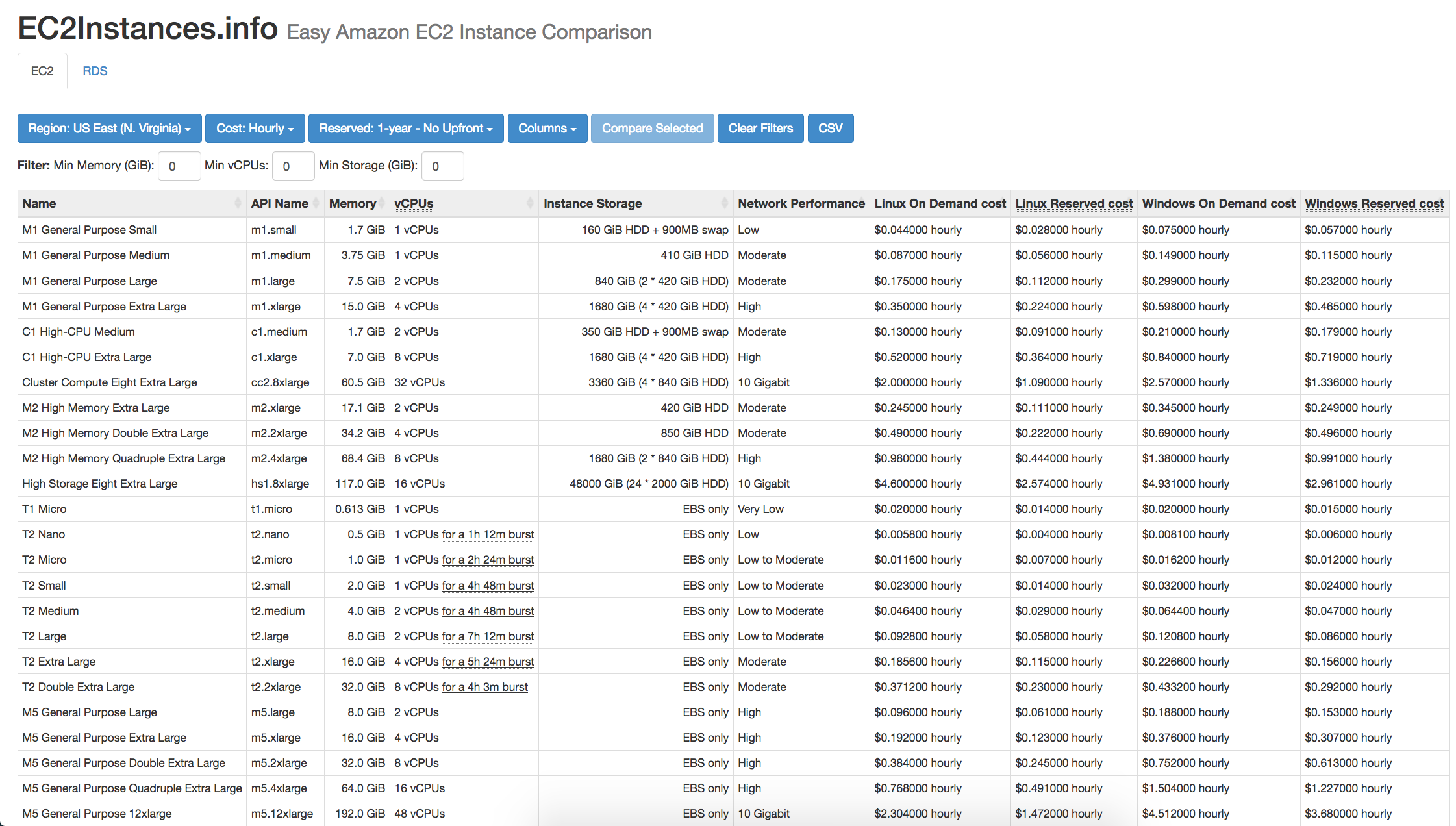
ec2instances.info
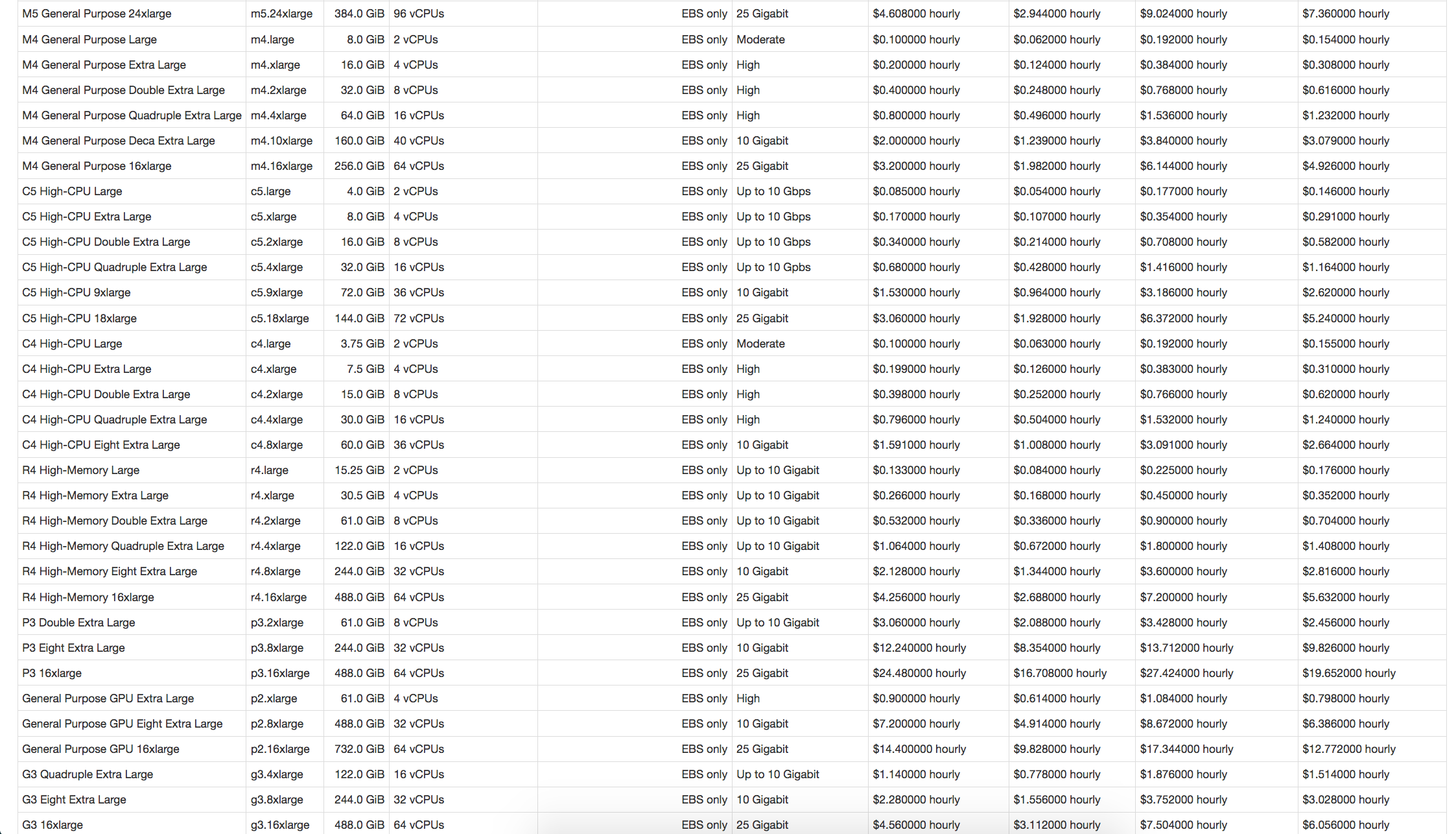
ec2instances.info
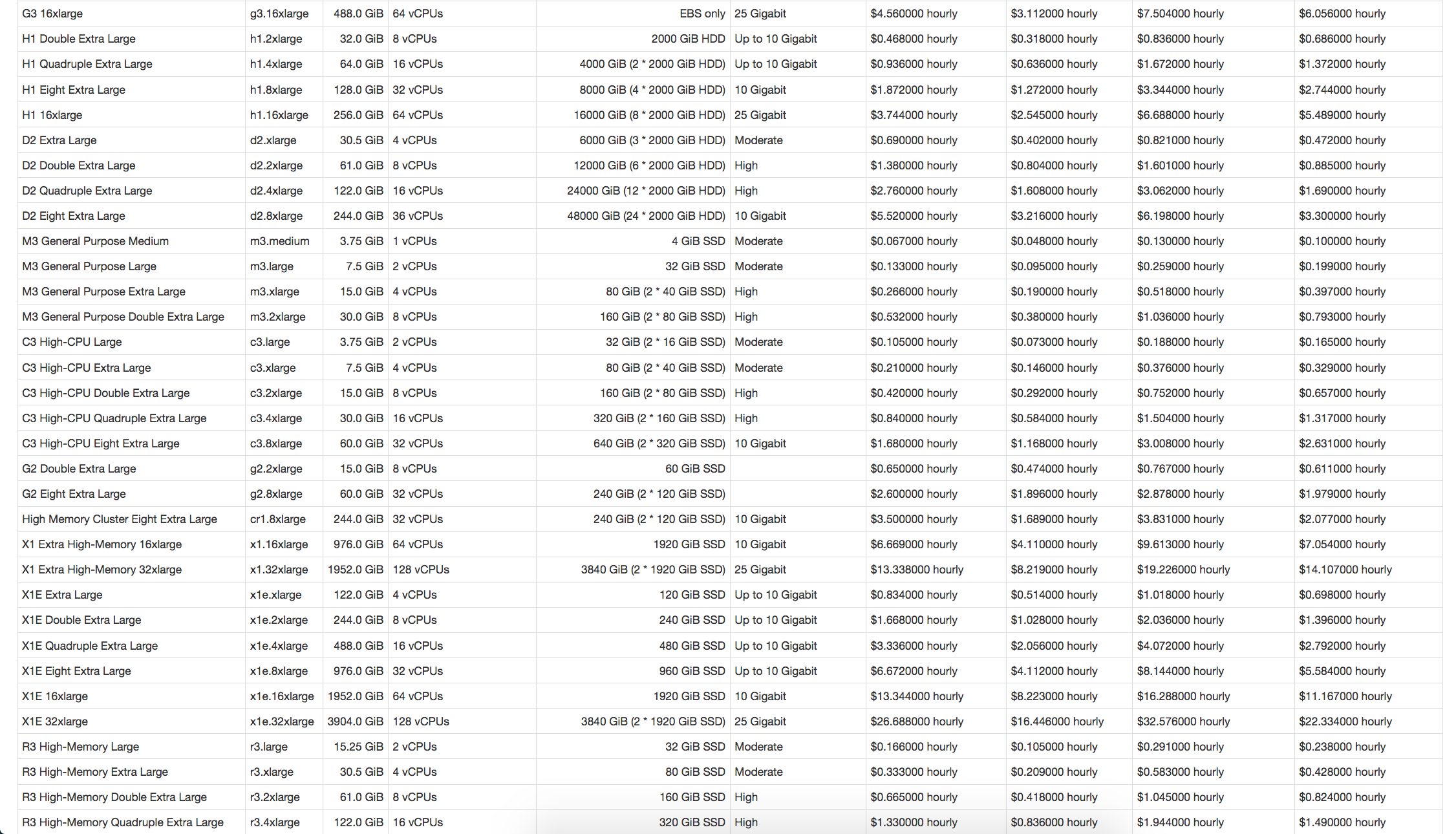
ec2instances.info
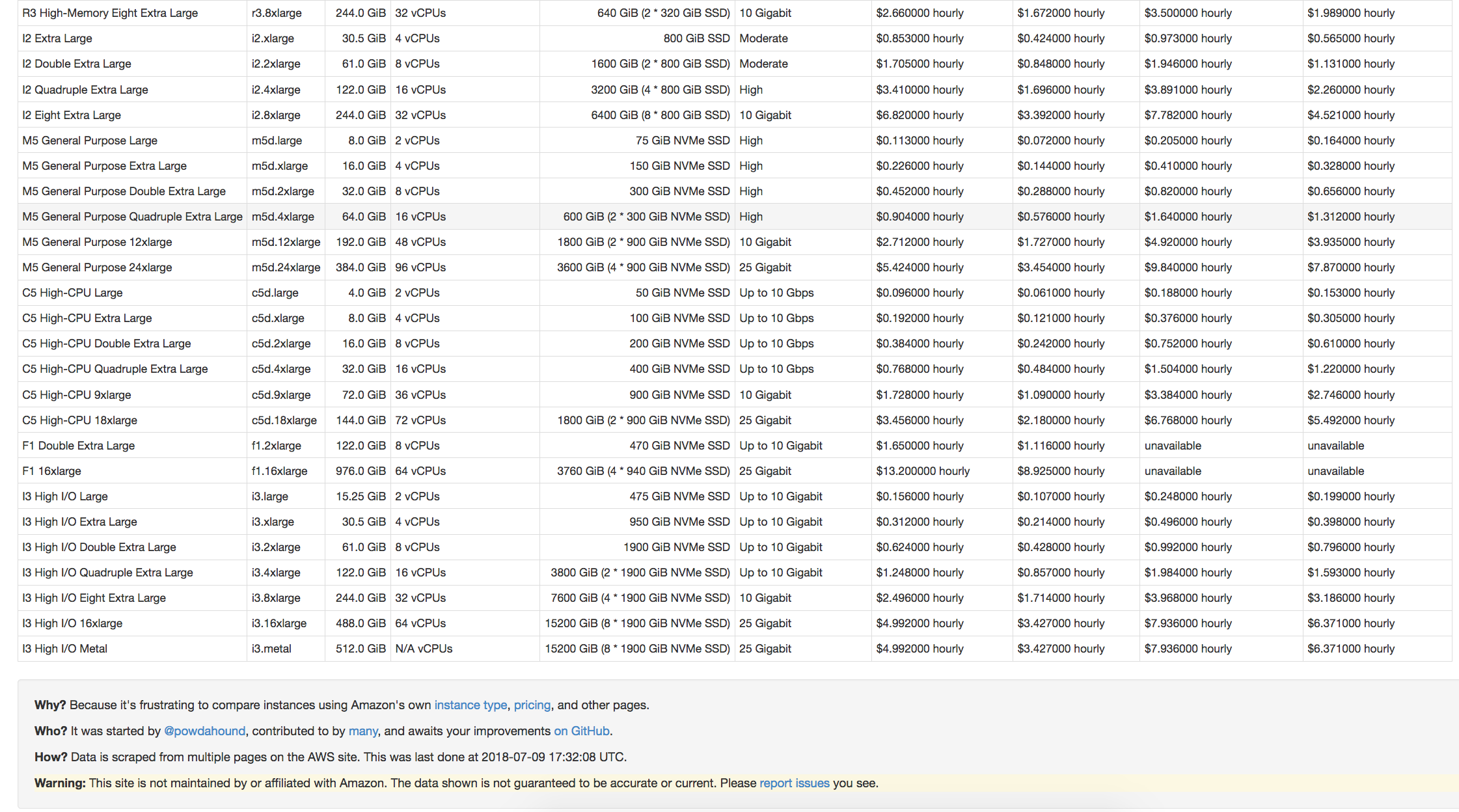
ec2instances.info
AWS Batch

Pros:
- Abstracts away infrastructure details
- Dynamically provisions AWS resources based on requirements of user-submitted jobs
- Allows scientists to run 100,000+ batch jobs
Cons:
- AWS Web Console resists automation
- Does not easily facilitate reproducibility
- Requires learning new terminology
AWS Batch workflow
- Build a Docker image (local machine)
- Create an Amazon ECR repository for the image (web)
- Push the build image to ECR (local machine)
- Create IAM Roles, compute environment, job queue (web)
- Create a job definition that uses the built image (web)
- Submit jobs (web)
Challenge
Reap the benefits of AWS Batch from the comfort of our Python env
Previous attempts
Other projects have sought to lower AWS barrier to entry
- PiCloud (2010), acquired by Dropbox in 2013
- pyWren (2017), built on AWS Lambda
- 5 minute execution time
- 1.5 GB of RAM
- 512 MB local storage
- no root access
Cloudknot
Single Program
import cloudknot as ck
def awesome_func(...):
...
knot = ck.Knot(func=awesome_func)
Multiple Data
import cloudknot as ck
def awesome_func(...):
...
knot = ck.Knot(func=awesome_func)
...
future = knot.map(args)
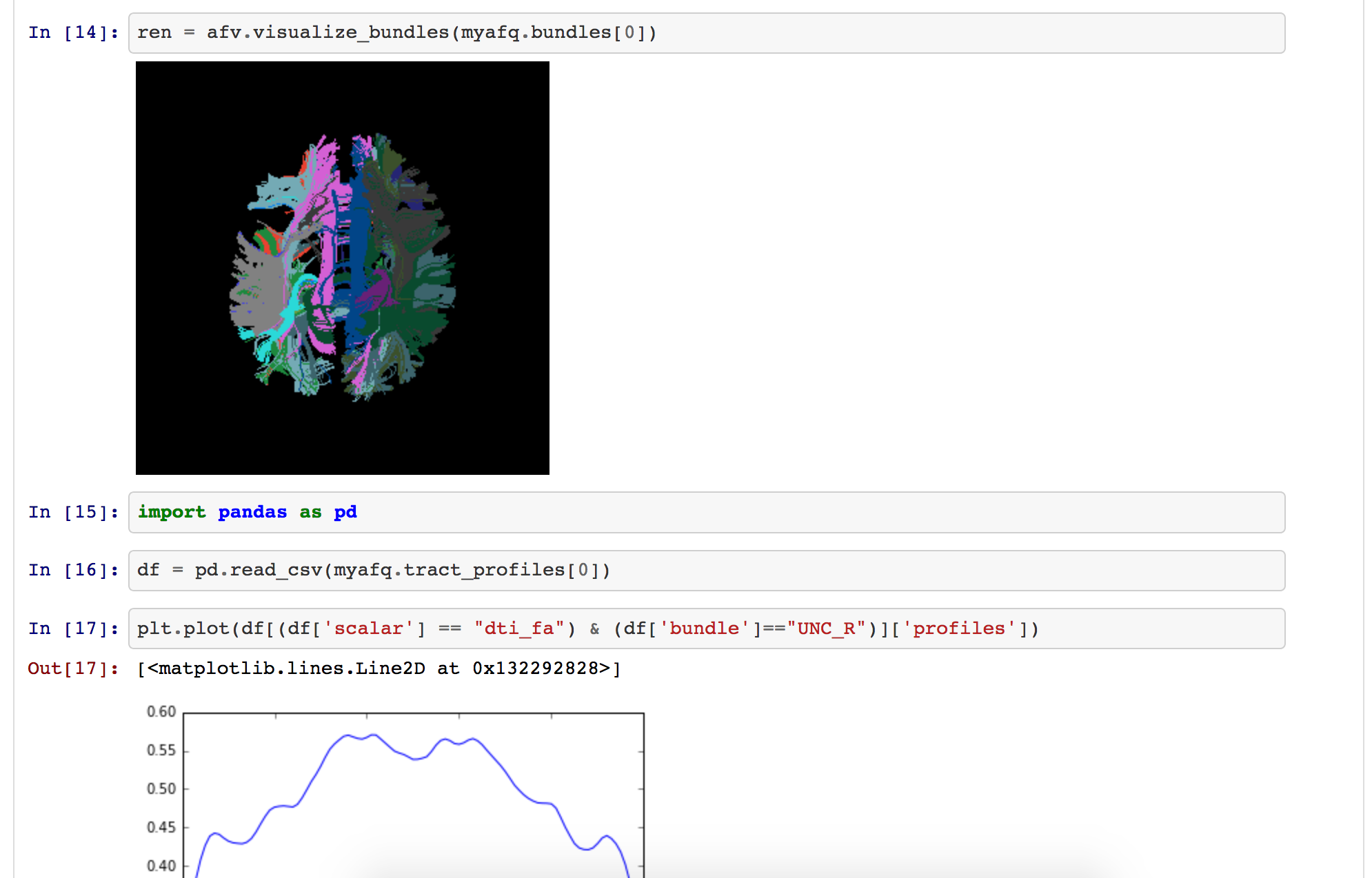
Analysis of MRI data using DIPY (Garyfallidis et al., 2014)
Brain extraction
Denoising
Tensor fitting
(see code)Compare to Dask, Myria, Spark using previous benchmark study (Mehta et al., 2017).
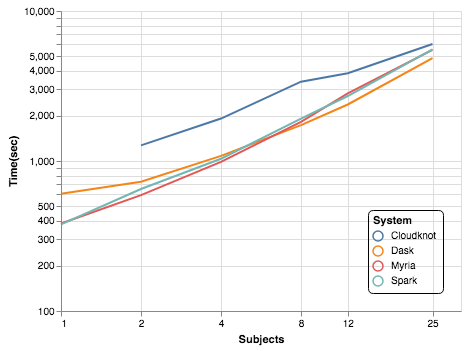
Analysis of MRI data
Takeaway
- Previous MRI benchmark was performed by a team of 4 graduate students and postdocs over 6 months.
- Cloudknot implementation took Ariels one day
- For 25 subjects, Cloudknot was 10-25% slower
- Cloudknot favors workloads where development time is more important than execution time
Conclusion
- Cloudknot favors workloads where development time matters more than execution time.
- For many data science problems, this is an acceptable trade.
- Simplified API makes cloud computing more accessible.
import cloudknot
knot = cloudknot.Knot()
results = knot.map(sequence)
Additional examples
Github repo: https://github.com/nrdg/cloudknot
Documentation: https://nrdg.github.io/cloudknot/index.html
We welcome issues and contributions!
Thanks!
Adam Richie-HalfordJohn Kruper
Eleftherios Garyfallidis (Indiana University)
NIBIB CRCNS grant 5R01EB027585-02