
Statistical learning of human brain structure
Ariel Rokem, University of Washington eScience Institute
Follow along at

Normal behavior is supported by brain connectivity
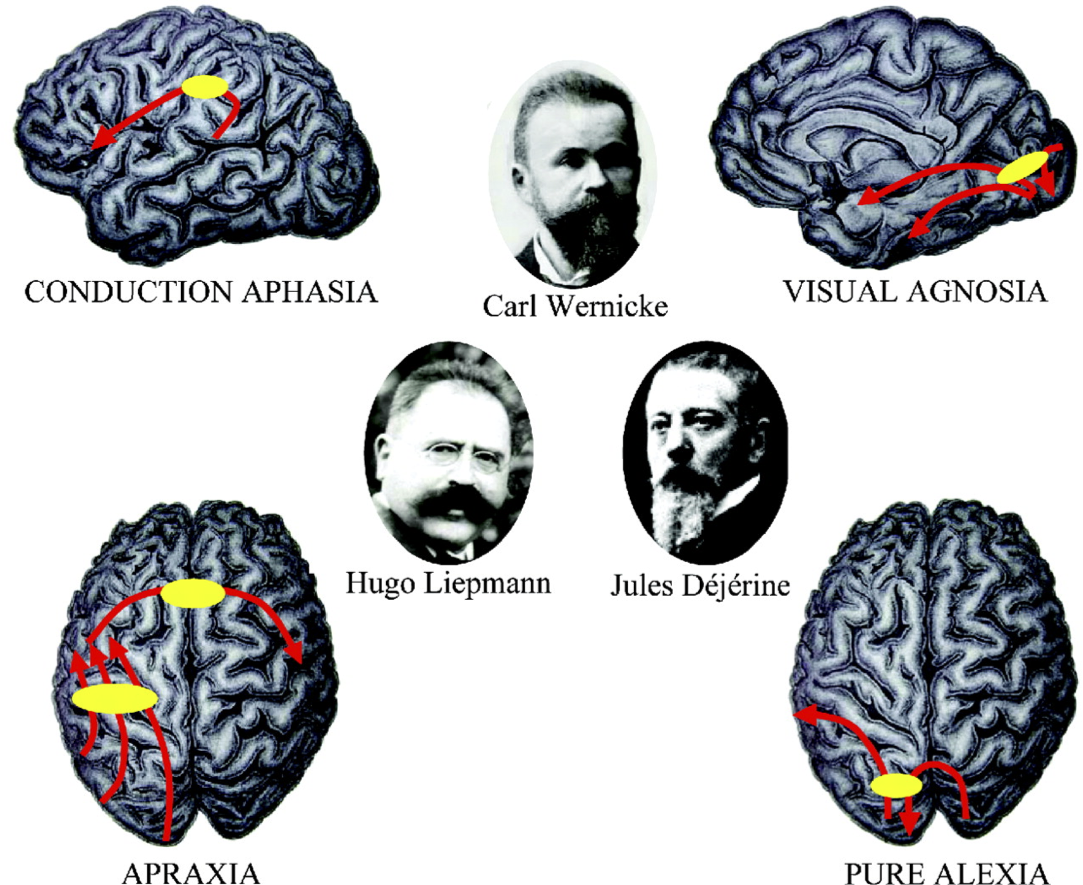
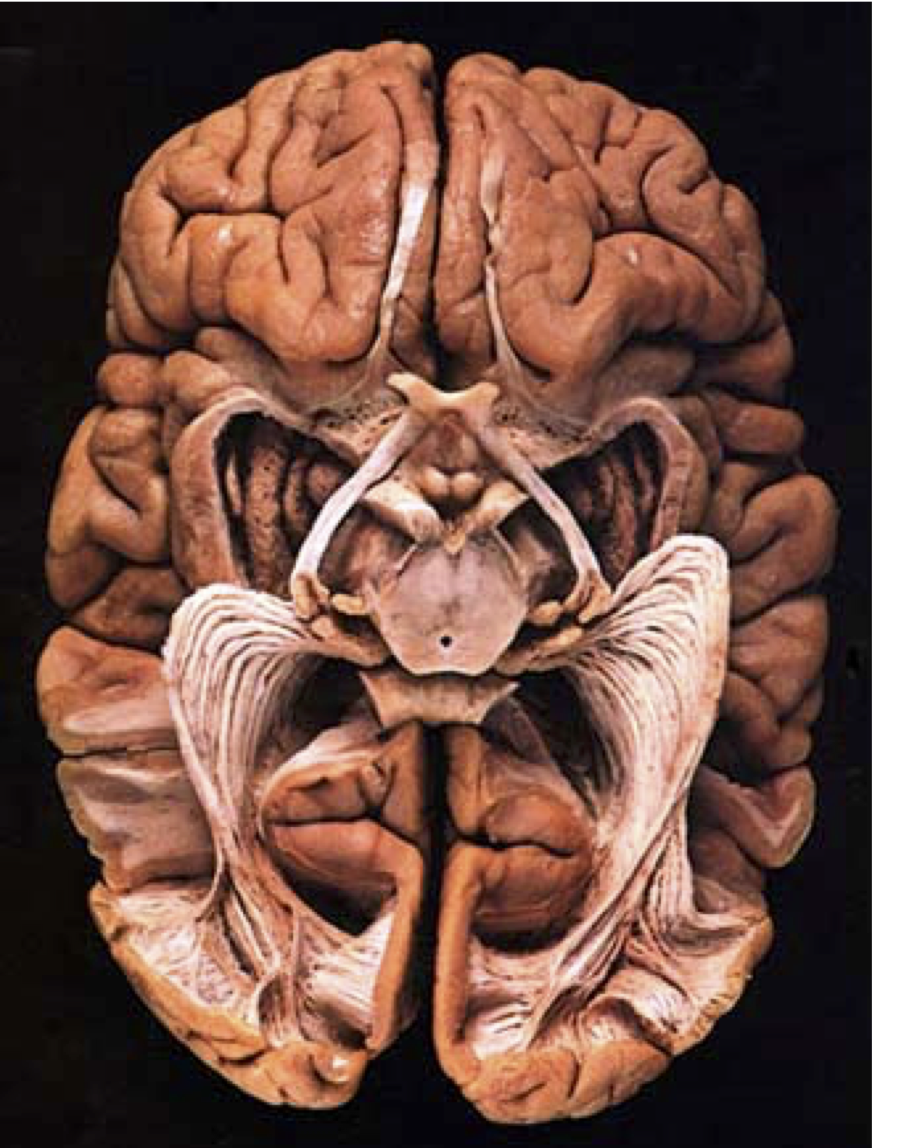
Not just passive cables
Brain connections change with development
Individual differences account for differences in behaviour
Adapt with learning
This has clinical significance
Magnetic Resonance Imaging (MRI)
Neural activity: functional MRI
Anatomy: structural MRI
...
Brain connectivity: diffusion MRI

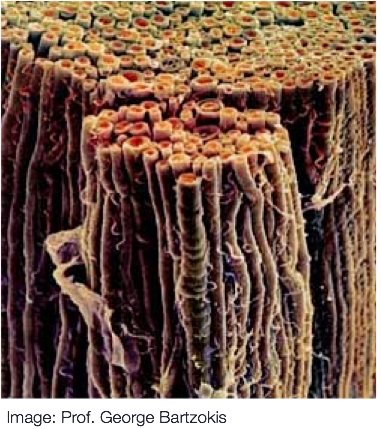
Diffusion MRI
Diffusion MRI


Diffusion MRI
Modeling diffusion
Diffusion statistics
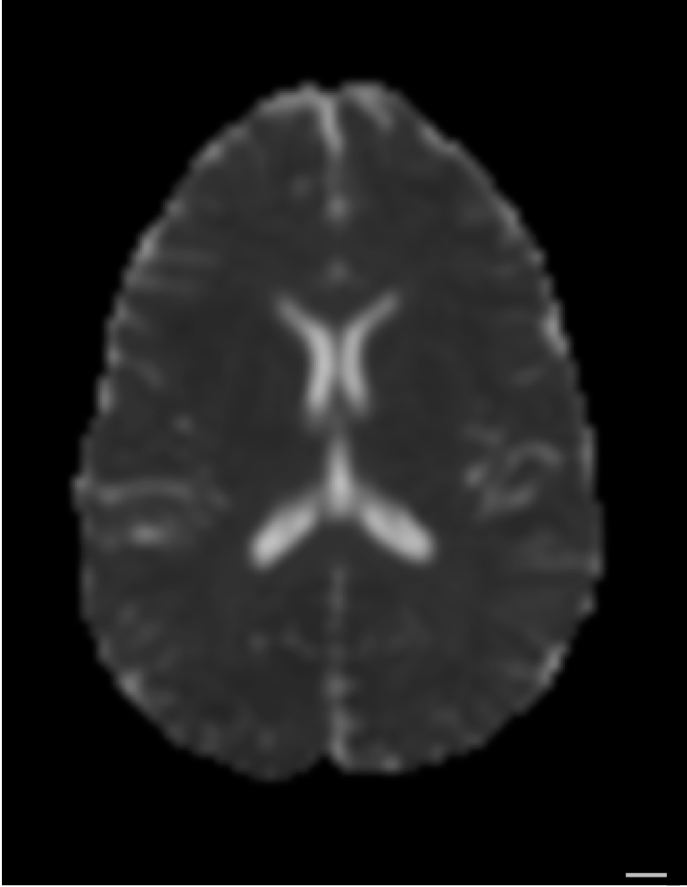
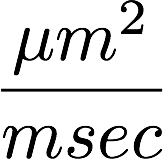
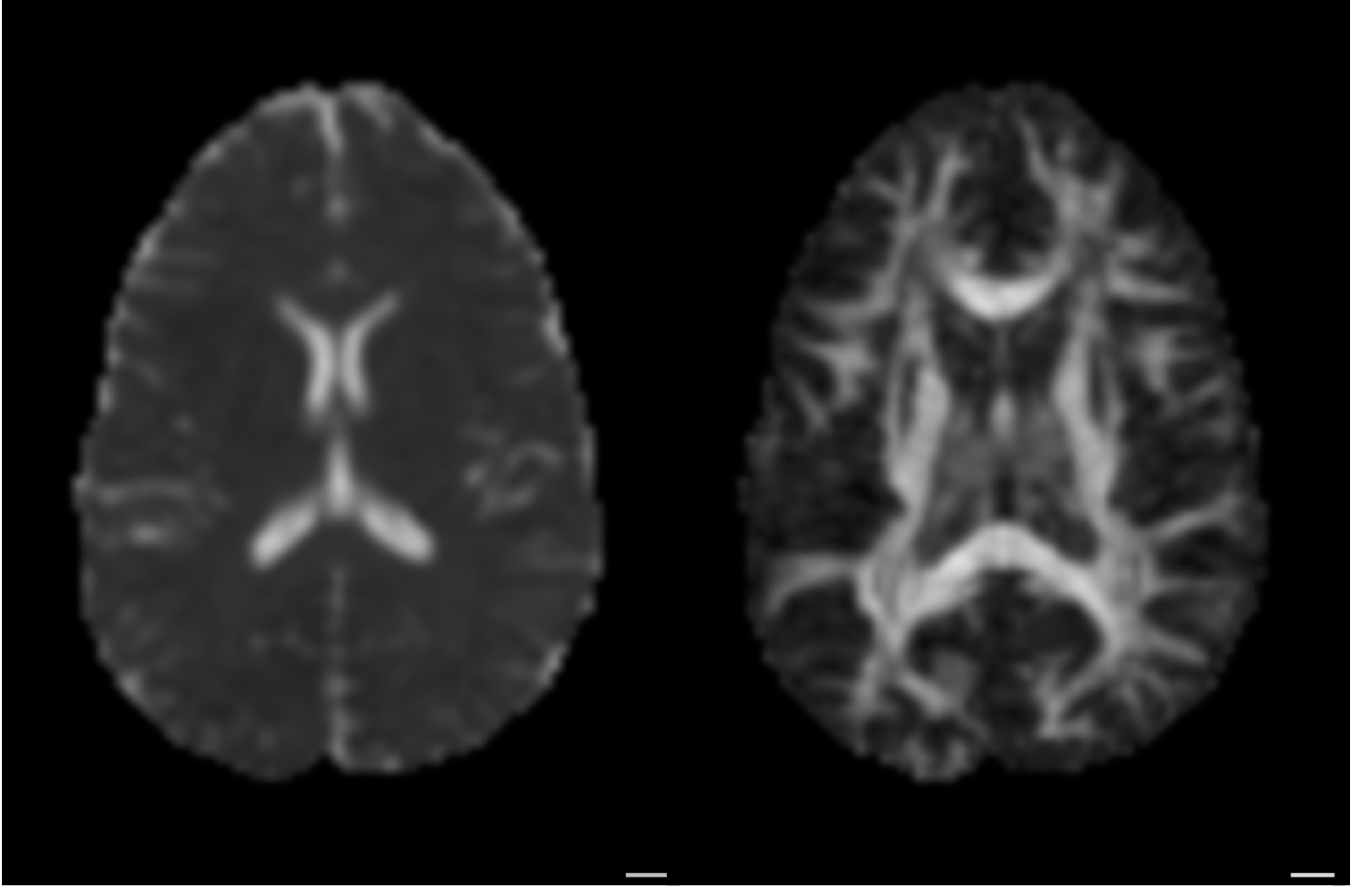
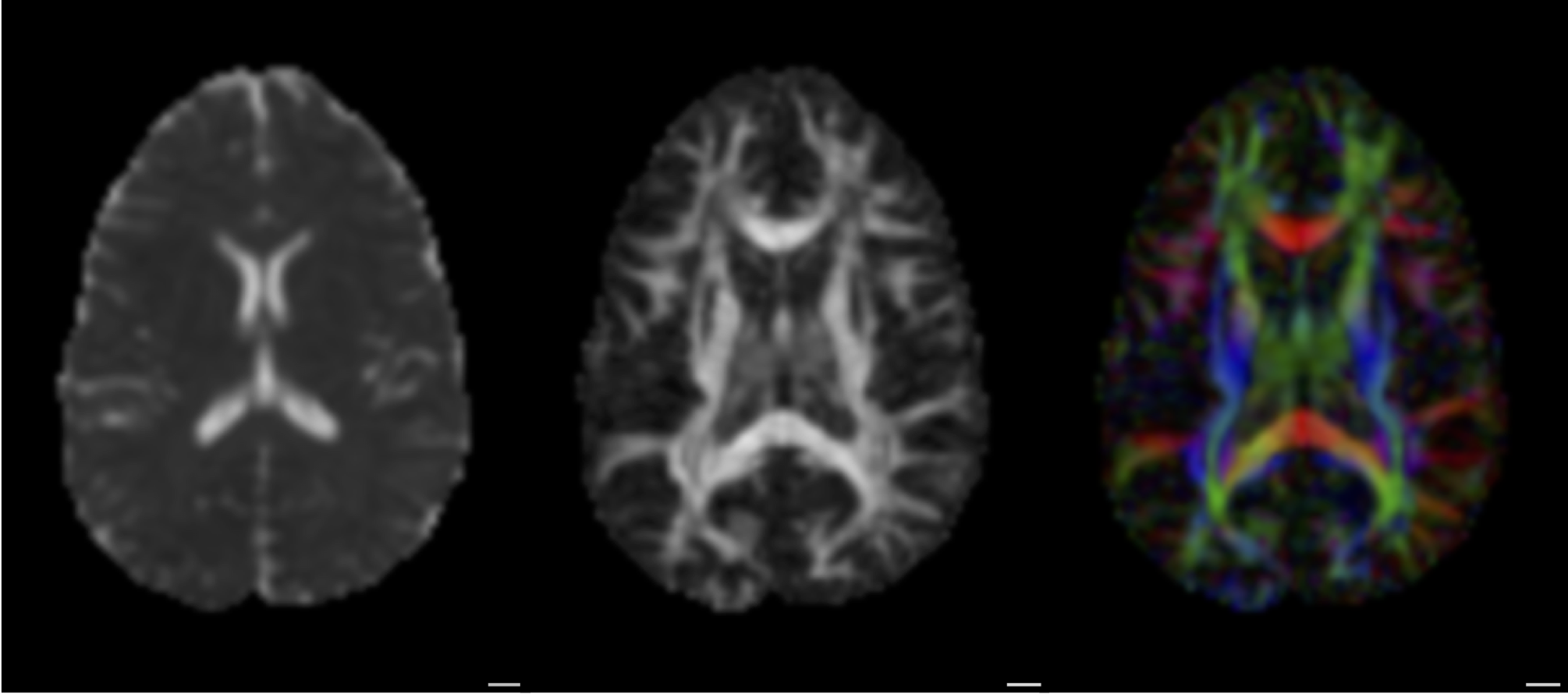
From diffusion to tracks
From diffusion to tracks


From diffusion to tracks
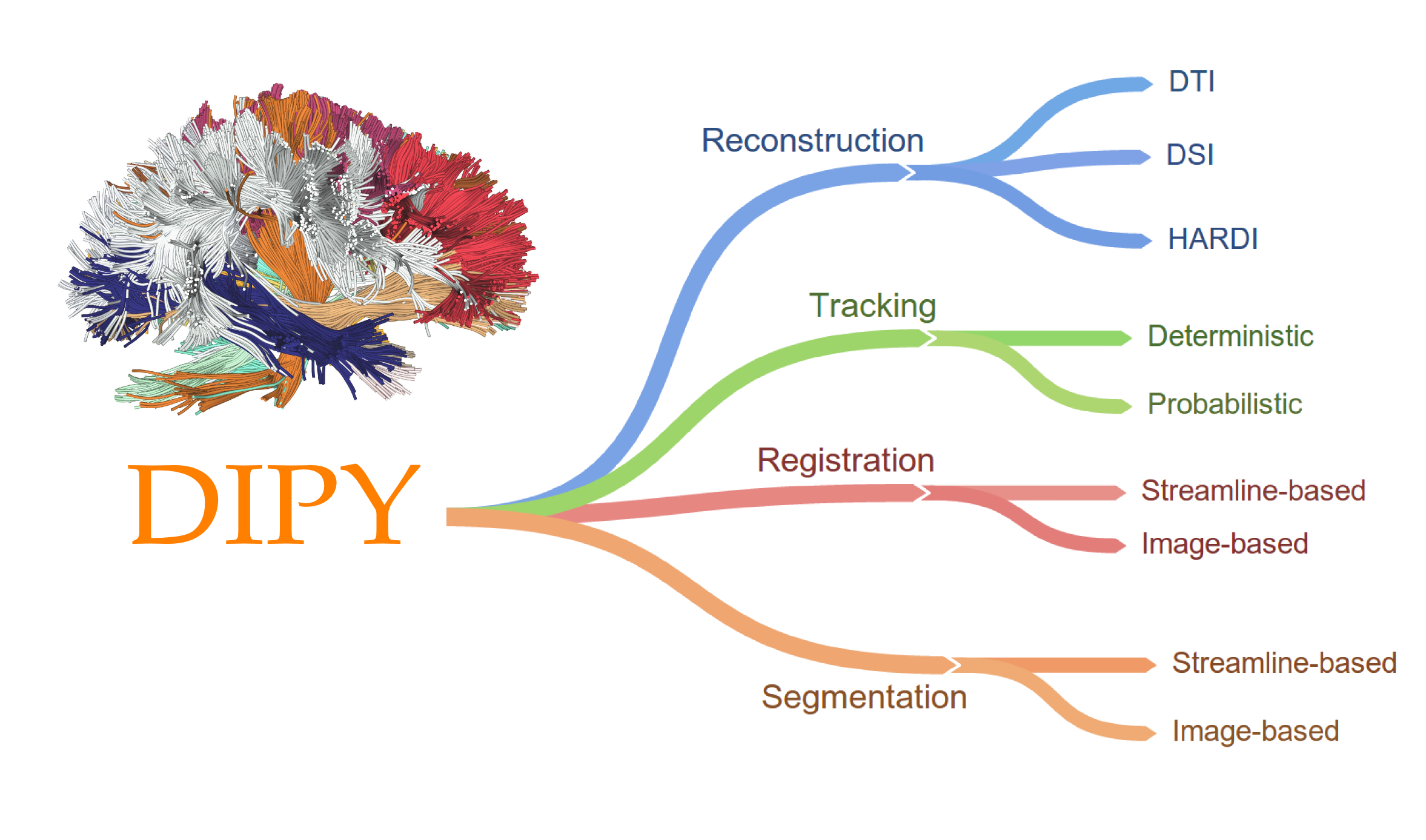
DIPY: Diffusion MRI in Python
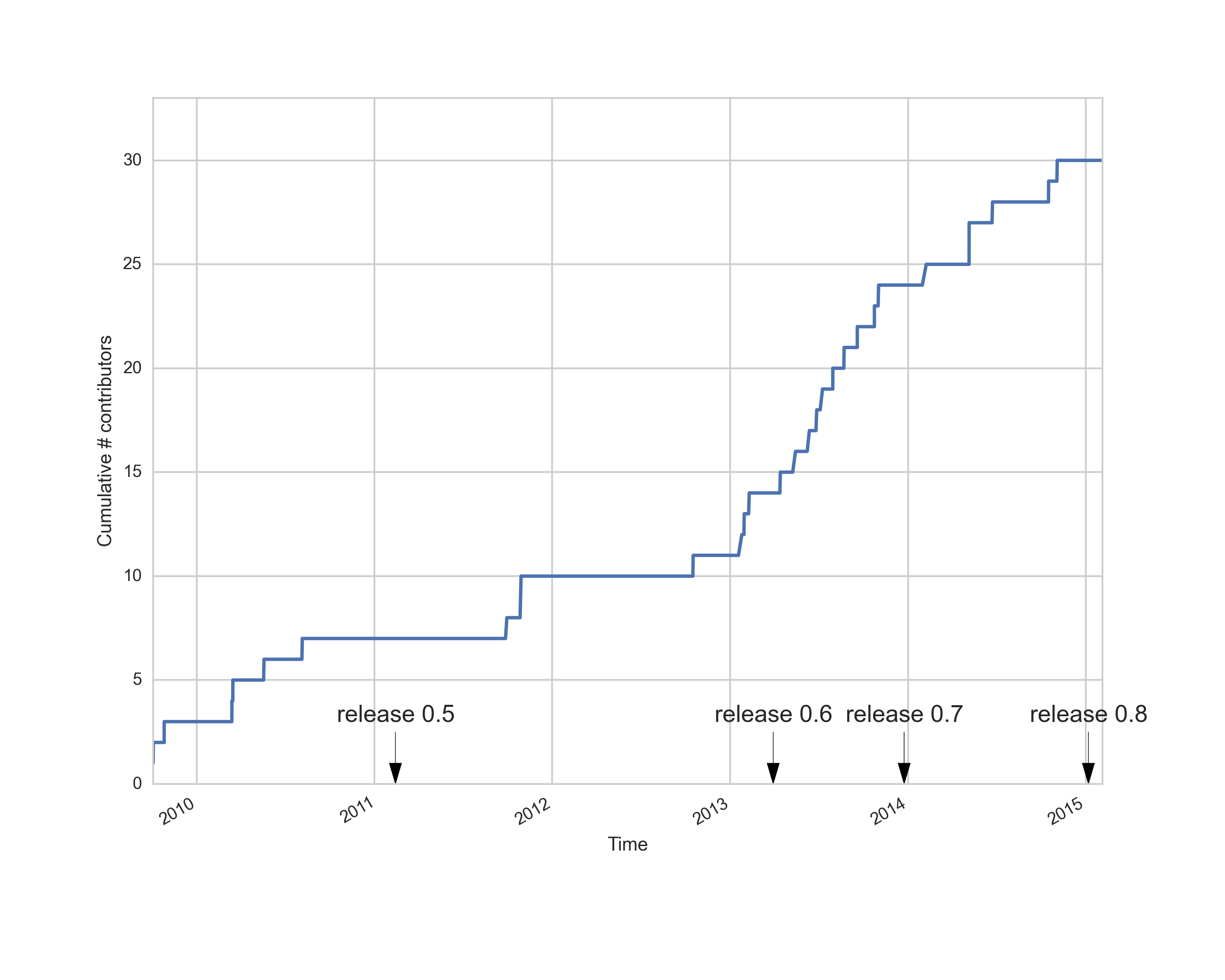
Started in 2009 by Eleftherios Garyfallidis
Contributors from at least six different countries and many different labs
Diffusion MRI: the challenge of validation
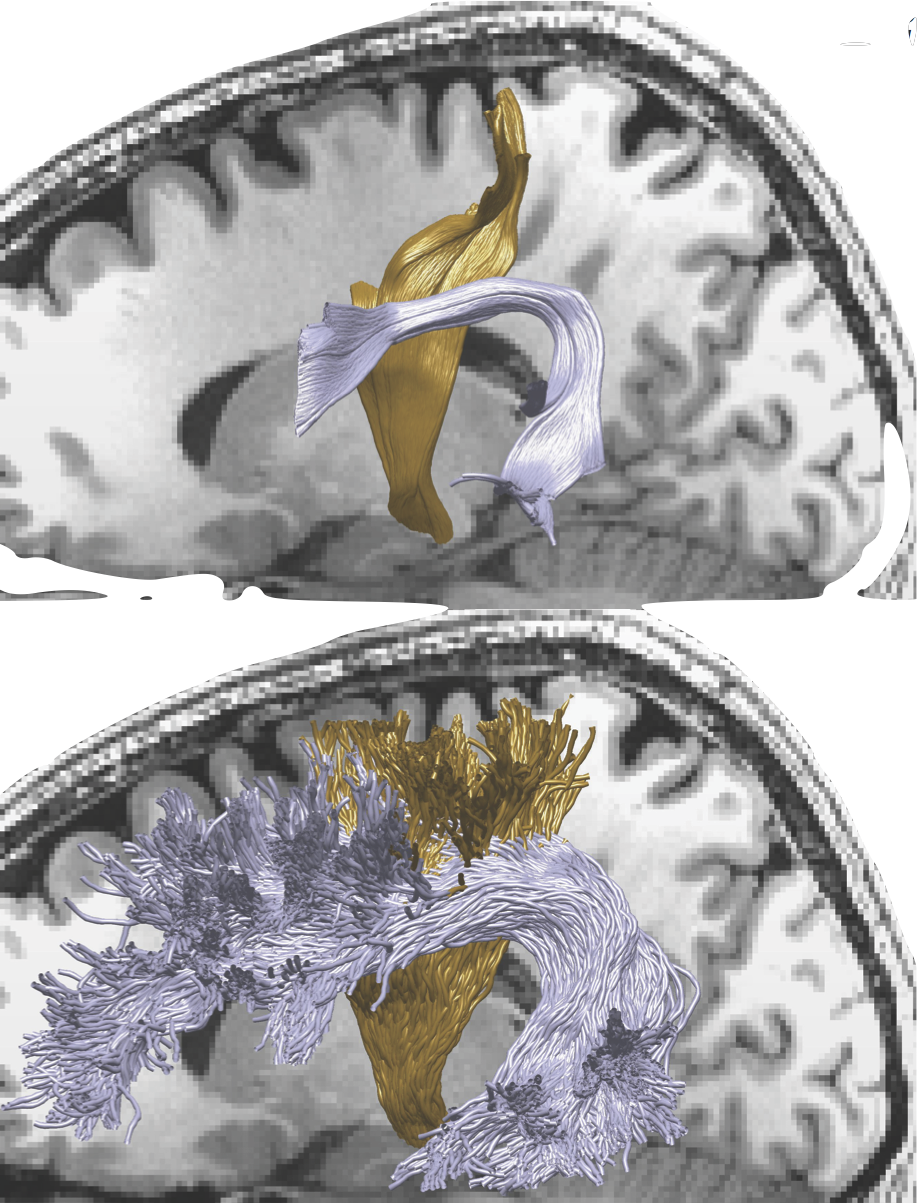
A statistical learning approach
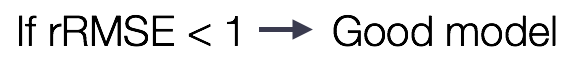


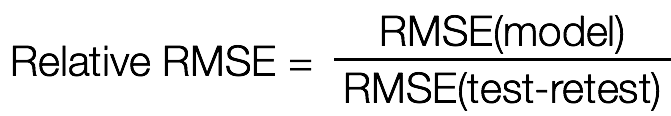

Dipy cross-validation API
gtab = gradient_table(...)
model = ReconstModel(gtab, ...)
fit = model.fit(data, ...) # => ReconstFit
prediction = fit.predict(gtab, ...)
For example
model = dti.TensorModel(gtab)
fit = model.fit(data1)
prediction = fit.predict(gtab)
RMSE = np.sqrt(\
np.mean((prediction - data2) ** 2), -1))
rRMSE = RMSE / np.sqrt(\
np.mean((data1 - data2) ** 2), -1))
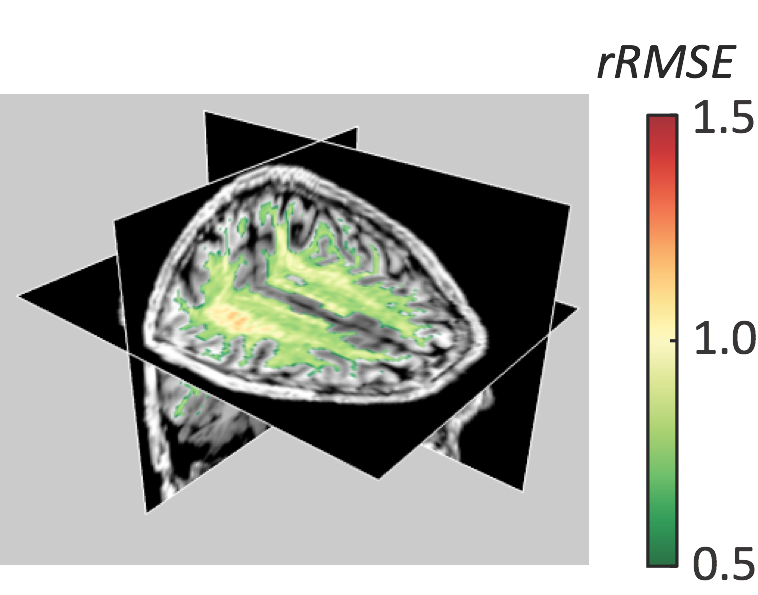 Rokem et al. (2015)
Rokem et al. (2015)
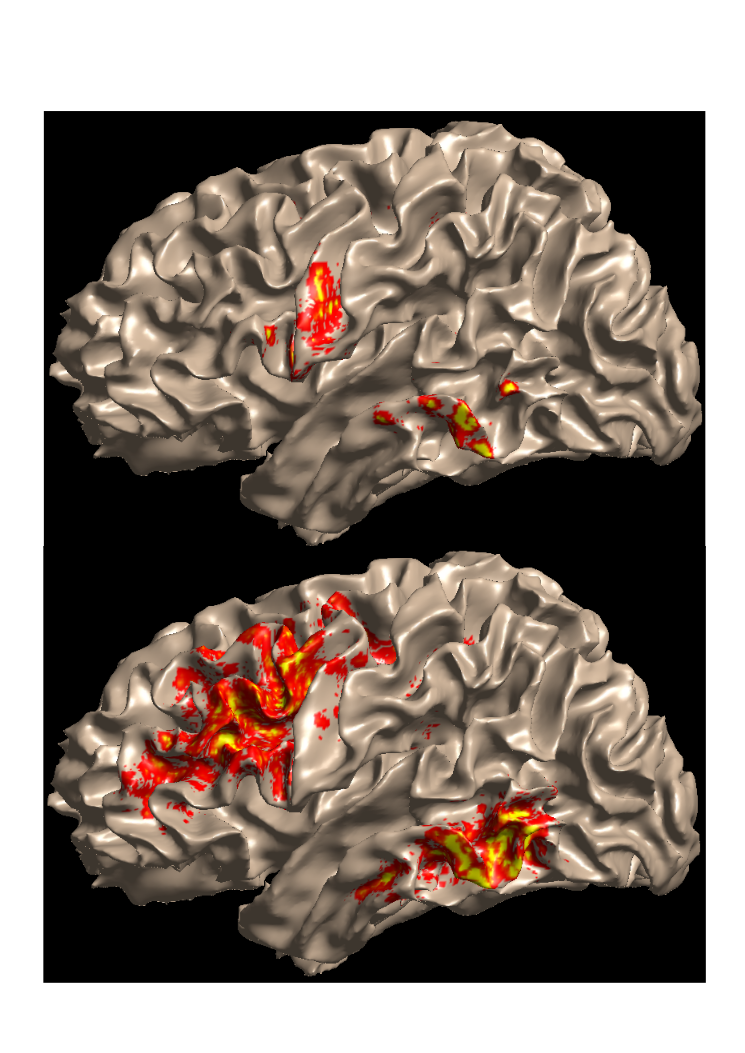
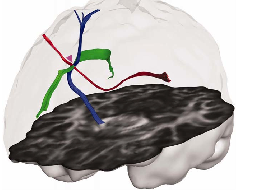 Corpus callosum
Corticospinal tract
Superior
Corpus callosum
Corticospinal tract
Superior
longitudinal fasciculus
DTI
 Crossing fiber model
Crossing fiber model
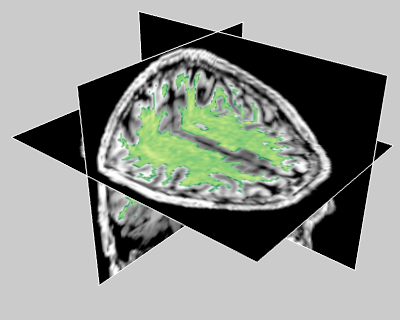 Rokem et al. (2015)
Rokem et al. (2015)
When you've only measured once
k-fold cross-validation
# Use a k of 2
dti_pred = kfold_xval(dti_model, data, 2)
csd_pred = kfold_xval(csd_model, data, 2)
 Algorithm 1
Algorithm 2
Algorithm 1
Algorithm 2
LiFE: Linear Fascicle Evaluation
Forward model from the tracks to the measured signal
Pestilli et al. (2014)
From diffusion to tracks


From tracks to diffusion
 ...
=
...
=
 Pestilli et al. (2014)
Pestilli et al. (2014)
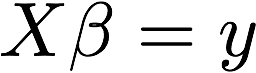 Solve for
Solve for
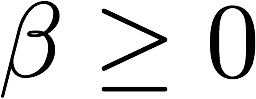
>>> X.shape
(10e8, 10e6)
Pestilli et al. (2014)
fiber_model = life.FiberModel(gtab)
fit = fiber_model.fit(data, tracks)
prediction = fit.predict(gtab)
optimized_tracks = tracks[fit.beta>0]
Summary
Measuring brain connectivity with diffusion MRI
The Dipy project
The validation problem
In vivo validation through statistical learning
Collaborators
Dipy:
 Eleftherios
Eleftherios
Garyfallidis
 Stefan
Stefan
Van der Walt
 Bago
Bago
Amirbekian
Collaborators
Stanford VISTA lab:
 Brian
Brian
Wandell
 Franco
Franco
Pestilli
 http://arokem.org
http://arokem.org
 arokem@gmail.com
arokem@gmail.com
 @arokem
@arokem
 github.com/arokem
github.com/arokem
